Canada supporting DNA-based biomonitoring for Oil Sands watersheds and more
Brand new DNA-based environmental monitoring methodologies are emerging, thanks to the pioneering work of a collaboration of Canadian genetics researchers and federal support.
The Canadian government plans on contributing $280,000 to the University of New Brunswick’s Canadian Rivers Institute, known for their expertise in watershed science and ecological sustainability, further develop gene-based monitoring method development.
The contribution is only the latest show of support for DNA-based monitoring methods in Canada, following a $3.2 million grant from Genome Canada. The government’s interest in the work comes from its strong desire to improve environmental monitoring of its watersheds all over the country. Water quality monitoring of the area of the Joint Oil Sands Monitoring Program has been a focus of the research, for example.
Mehrdad Hajibabaei, assistant professor of integrative biology at the University of Guelph, is the chief developer of the new DNA-based methods. Hajibabaei works closely with Donald Baird, CRI science director, who aids in application of the DNA sequencing methodologies for use in the institute’s effort for biodiversity and ecological risk assessment.
According to one of Hajibabaei and Baird’s publications in PLOS ONE, the genetic marker methods used involve selection of specific genes or DNA sequences from an organism’s genome. DNA sequencing information from these short sequences, or “barcodes,” can identify organisms down to species.

Comparitive analysis of environmental DNA barcodes. (Credit: Ontario Genomics Institute)
Hajibabaei and Baird’s DNA methods can identify organisms at the species level in days instead of the weeks or months required for traditional processing and reporting. Baird has done extensive work on freshwater benthic macroinvertebrates, though the DNA sequencing could be used for any living creature of interest with a known genetic library.
Scientists would traditionally identify organisms one by one based on their morphology — an accurate but time-consuming method. Typically environmental samples containing hundreds to thousands of individual organisms must be processed in the lab. New DNA sequencing technologies can now be performed on bulk environmental samples without the need for sorting and many samples can be analyzed simultaneously. Additionally, a single sample can yield much more information.
Referred to generally as the Biomonitoring 2.0 project, the new DNA-based environmental monitoring approach includes the most advanced sample gathering techniques, genomic sequencing methods, bioinformatics and methods for data dissemination and integration. The new methodology uses genetic markers for monitoring ecosystem health and is expected to be less expensive and more efficient than current monitoring methods.
The genetic marker methods are a big advancement in monitoring brought about by a joint effort among the University of Guelph, Environment Canada and the Canadian Rivers Institute with financial support provided by Genome Canada. The Biodiversity Institute of Ontario of the University of Guelph is a key participant in support of method development.
Biomonitoring 2.0 will be integrated in Canadian Aquatic Biomonitoring Network Protocols. Through use of CABIN protocols and training, data will be collected in a rigorous and consistent manner throughout Canada and will be suitable for use by academics, industrialists, consultants, volunteers, First Nations, and provincial and territorial researchers.
Michelle Gray, co-investigator and director of training and program development of UNB-CRI, says in a recent UNB blog entry that the new genetic marker methodology will help CRI realize its “core objective of merging research science with environmental and society needs.”
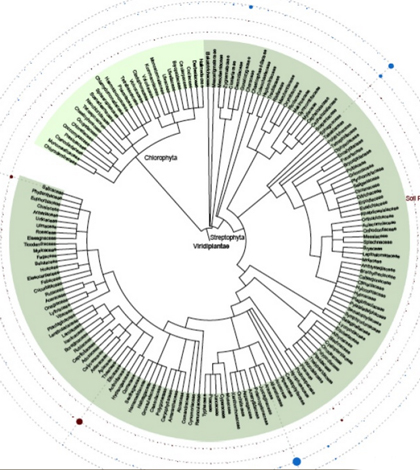
Top image: Phylogenetic profile and diversity of plants and algae. (Credit: Hajibabaei et al)











0 comments